Herbert Spencer would be so proud.
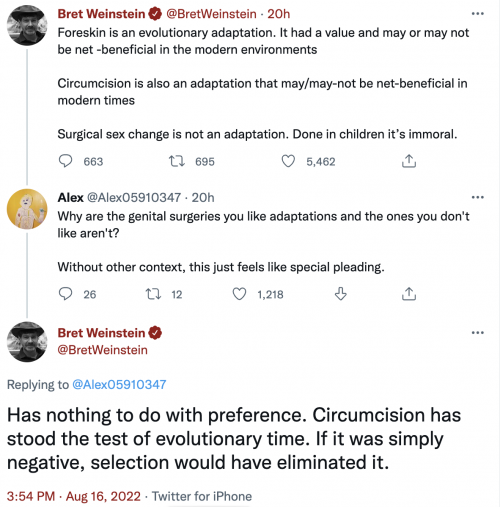
One quibble: evolution by natural selection isn’t the fundamental concept, it’s evolution, change, by multiple mechanisms.
The red button comic is also relevant.
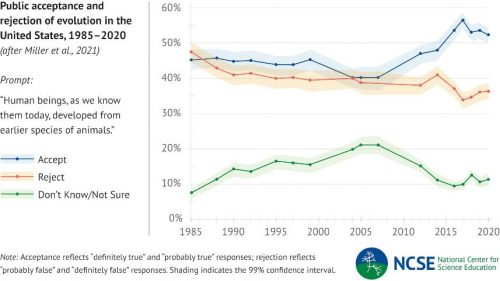
Yeah, because it’s virtually impossible to teach a creationist anything.