I have a bit of a peeve with a common analogy for the human genome: that it is the blueprint of the body, and that we can find a mapping of genes to details of our morphological organization. It’s annoying because even respectable institutions, like the National Human Genome Research Institute, use it as a shortcut in public relations material. And it is so wrong.
There is no blueprint, no map. That’s not how the system works. What you actually find in the genome are coding genes that produce proteins, coupled to regulatory elements that switch the coding genes off and on using a kind of sophisticated boolean logic. Each cell carries this complex collection of regulated genes independently and identically, but the boolean logic circuits produce different outputs varying with the inputs from the environment and the diverging histories of each cell. For instance, there is no code anywhere in the genome that commands the forelimbs to make five and only five digits: instead, a cascade of genes and cell movements produce a patterned tissue that in us contains sufficient mass and is of a size to generate five nuclei of condensing tissue that produce fingers.
It’s better to think in terms of cellular automata. The embryo is a pool of autonomous cellular robots that have general rules for how they should respond to environmental cues…and those cues tend to vary in predictable ways across the embryo, leading to a consistent cascade of action that produces a relatively consistent complex product, the multicellular organism.
The unfortunate consequence of those properties, though, is that you’ll never be able to look at a single gene from the genome and sort out what it does in the embryo. All the genes will be rather cryptic; you might be able to figure out that, for instance, the gene codes for an adhesion protein that makes the cell stick to a certain other class of cell, and that it’s switched on by gene products X and Y and turned off by gene product Z, but obviously you won’t be able to figure out its role until you figure out what activates genes X, Y, and Z, and whether the cell happens to be in a particular adhesive environment. And then when you look at X, Y, and Z, you discover that they have similar patterns of conditional logic in their expression.
In order to understand what a particular gene does, you have to understand what all the other genes do, as well as all the details of signaling and cell interactions that are going on, oh, and also, it’s entire developmental history, since epigenetic interactions can shape the future behavior of a cell lineage.
Hey, let’s all give up. This stuff is too hard.
No, let’s not. What it means is that you can’t derive the organism from the mere sequence of the genome — that is, the genomic information is not sufficient to comprehend morphology, because developmental processes add extra-genomic information to the generation of the organism. It means developmental biologists have job security (yay!), because the only way to decipher what is going on is to work backwards, from morphogenetic/physiological events to the underlying genes involved. This is not to imply that the genomic information is unimportant, only that understanding it requires complementing it with an understanding of cell:cell interactions, signaling, signal transduction, induction, and molecular patterning…all stuff that developmental biologists love.
Now if all you get from this is that the genome and organism are complex, interlocking, interdependent features that are so immensely and tightly integrated that evolution must be impossible, you aren’t thinking like a developmental biologist yet. Ask an evo-devo person about this stuff, and they’ll tell you that this is great…the way development works makes evolution of form easy.
That’s because there is no blueprint. What you have instead is a collection of flexible robots that have this property called plasticity: give them a novel environment or condition, and they don’t curl up and die and do nothing. Instead, they just follow the rules they’ve got and try to make something coherent out of whatever situation they find themselves in. They aren’t committed to making five fingers in any way; give them a reduced tissue mass, or an enlarged mass, or a variation in the signaling environment, and they’ll build something. And often it’s something surprising. Development is really, really good at producing emergent properties, precisely because it is autonomously rule-based rather than blueprinty.
All this buildup has a point: there’s an evolutionary issue that has a developmental resolution. It’s some really cool work on the development of the limb.
So I work on zebrafish. They don’t have limbs, obviously, but they do have fins where we tetrapods build legs and arms. Fins are thin membranous folds of ectoderm (our fancy word for skin), infiltrated with thin rods of cartilage called fin rays. Developmentally, they arise from things called fin folds — flaps of ectoderm that flatten to form a double-walled sheet.
In development, tetrapods add an extra element to the fin fold: mesoderm, that tissue that forms bone and muscle, expands to fill the fin fold with the raw material of a muscular, bony limb.
You can visualize the developing limb as something like a slab of pita pocket bread. Fish are content with just the bread, giving it a little reinforcement, while tetrapods open up that hollow space and stuff it full of filling. That filling represents a field of great potential, which is then organized in reproducible ways to make limb bones and digits and muscle. The question addressed here isn’t about the precise organization of the limb, but a more general one about where all that tasty filling came from in evolution.
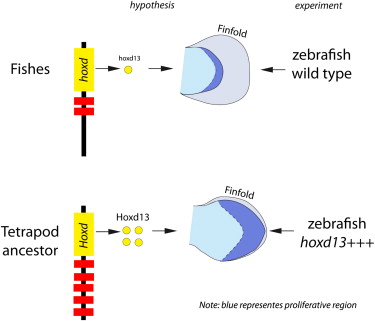
We have hints. There are genes switched on in the distal part of the fin/limb that are more strongly activated in tetrapods than in fish; these genes are associated in space and time with an increase in the volume of mesodermal tissues. The gene of interest here is called Hoxd13. It’s one of a well-known array of genes that are responsible for patterning the body axis, some of which have also been recruited into patterning the limb. The hypothesis is that expressing greater levels of Hoxd13 in a fish fin would lead to an expansion of mesoderm that would be a potential evolutionary precursor to turning a fin into a leg.
So here’s what Freitas and others did, and this just blows me away that we can do these kinds of transgenic experiments routinely nowadays. They made a construct of a Hoxd13 gene coupled to a glucocorticoid switch: just by exposing the developing fish to dexamethasone (Dex), they can turn the gene on. It’s like adding a volume control to a gene that they can turn up at will. They also used other techniques, coupling Hoxd13 to a heat shock promoter, so they could also turn it on just by putting the embryos in warmer water. It’s power. We can have complete control of a gene, and ask what happens when we overexpress it in a fish.
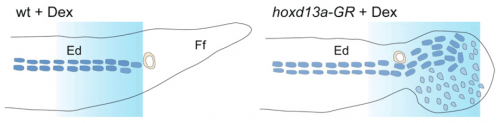
When you activate Hoxd13 at an appropriate stage in fin development, here’s a diagrammatic illustration of the results:
“Ed” is the endoskeletal disc; it’s the mass of mesodermal tissue that is found at the base of the fish fin, and that fills the whole tetrapod limb. “Ff” is the fin fold, the ectodermal flap that makes up the fin. “Ff” is the pita bread, “Ed” is the filling.
Switching on an excess of Hoxd13 has a couple of effects. One is that another gene, cyclin d1 is also turned on at a higher level. The cyclins are cell cycle regulators; amping up this gene leads to greater proliferation, so more mesoderm is made in this region. This mass then floods into the fin fold, building a lumpy meaty mass that does the poor zebrafish no good, but looks like the core of a limb.
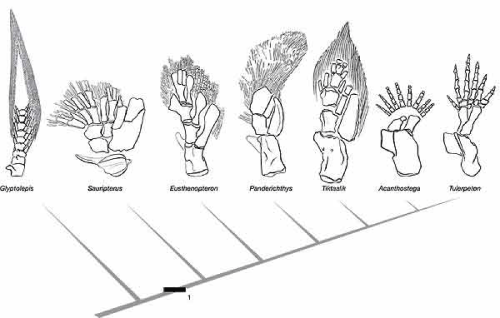
The fish does not build a hand or digits; it lacks the rules to carry out that degree of differentiation. But look at the limbs of these fossils from the fish-tetrapod transition.
There’s a lot of anatomical exploration going on in this series. This fits a model in which tetrapod ancestors carried a genetic variation that expanded the core of mesodermal tissue in their fins, which was then organized by the standard rules of limb mesoderm into bone and muscle. Again, this is opportunity, a new field of potential that in these early stages of evolution hadn’t yet been refined into a specific, and now familiar, pattern, although elements of that pattern are foreshadowed here.
This morphology fits a simple developmental model. The ancestral change was nothing more than the addition of new regulatory enhancers (and they have a candidate, called CsC, which is found in mouse but not zebrafish) that increased the expression of Hoxd13, which in turn led to an expansion of the raw material of limb mesoderm, which was then shaped by existing developmental rules into a crude bony, muscular strut.
Subsequent evolution refined that structure into a more specific limb morphology by layering new rules and new patterning elements onto the existing framework of genetic regulators.
So how did fish get legs? By progressive expansion of tissue that was then used autonomously by existing genetic programs to form a coherent structure, and which was then sculpted by chance and selection into the more familiar and more consistent shape of the tetrapod limb. Add raw material first, and the plasticity of developmental rules means that the organism will make sense of it.
The details are complicated, but complexity enables emergent evolutionary novelties. And that’s something beautiful about evolution and development.
Freitas R, Gómez-Marín C, Wilson JM, Casares F, Gómez-Skarmeta JL. (2012) Hoxd13 contribution to the evolution of vertebrate appendages. Dev Cell. 23(6):1219-29. doi: 10.1016/j.devcel.2012.10.015.



Or, if you have to use simple words (a lot of people who would understand the word “blueprint” will give you a blank stare if you say “cellular automata”), the metaphor of a “recipe” is not too bad. It has it’s problems, but as far as lay communication goes, it’s a pretty good metaphor. You follow the instructions, you get the intended structure.
No. IT’S NOT A RECIPE, EITHER.
That’s as misleading as the blueprint metaphor. There is no cook. There is no central description of the whole.
Posts like this why I keep coming back to Pharyngula. As it happens, just yesterday I got into a discussion with a friend about the “genome is a blueprint” idea (we were talking about the recent story of the potential to cloning a Neanderthal). I was trying to explain that the blueprint analogy is often used in the lay press (you know what I mean) because the reality is more difficult to express in limited column inches but that it, like all analogies, is a poor reflection of reality. This post puts it way better than I did. Thanks, PZ. Gonna share this.
FTR, I am an immunologist not a developmental biologist.
” What you have instead is a collection of flexible robots that have this property called plasticity: give them a novel environment or condition, and they don’t curl up and die and do nothing. ”
Well, within very narrow limits. You need the genome inside a zygote inside a womb or an egg, which in turn has to be within a healthy, unpoisoned female or an appropriate environment. It’s very easy to kill the thing. Most human zygotes never become babies, after all.
A bit of a quibble I suppose but very important to the story, no?
See, I think that is a narrow-minded idea of a “recipe” that, unfortunately, a lot of home cooks suffer from as well. A “recipe” in my mind is an outline of a process, and that can be varied in ways that are not “centrally described” in any way.
Take bread. I can take my basic recipe for bread, and by shaping the loaf in different ways and putting different cuts in the top, you get different shapes and textures. The dough is “programmed” to respond in certain ways by the whole process of kneading and rising, etc., but how it plays out is determined by environment and by other changes in the “recipe”. There is no dough recipe that says “Now be shaped like sandwich bread” or “now be shaped like a ciabatta”. The recipe just says, “Make a substance that will expand in these particular ways when heated, and will respond to neighboring dough/air in these ways.” Then the environment (the oven temperature, the cuts in the top, etc.) determine how it will actually develop.
I sort of get your point about there being no “cook”, but I challenge you to come up with a better metaphor that can be communicated to someone who doesn’t know the word “automata” :p I guess it’s like a recipe where each ingredient has a set of instructions for what to do with itself… But still, if the absolutely friendliest metaphor you are willing to put out there is “cellular automata”, you’re going to have a communication problem…
I also like the recipe metaphor because it captures the resilience of the process. If I am making chili, and instead of cooking the onions in some sort of fat before I add the liquid, I just throw uncooked diced onion in the liquid, it’s not like what I get is “no chili”, or something not even resembling chili, or a chili with no onions. It’s still a chili. A recipe can be modified, steps omitted, and you still get the same basic whole. There is no one-to-one correspondence between “steps” and “parts”, and changes in one step can effect multiple parts or the entire whole in non-linear ways.
Do you cook, PZ? :p
Which is why I said at the outset that while the Human Genome Project was not much ado about nothing; it was much ado about not all that much.
If we are looking for a metaphor that is absolutely perfect in every way, how about “A genome is like a genome” :p THAT’S helpful…
That is soooooo cool.
Biology rocks. Developmental biology diamonds.
Right, so the DNA is more like a bitmap encoding of a picture of the final organism.
bpcross wrote; “Which is why I said at the outset that while the Human Genome Project was not much ado about nothing; it was much ado about not all that much.”
Seriously? I don’t think anyone involved in the HGP or any other scientist trained in biology for that matter, thought that the project would give us a “blueprint” for an organism for exactly the reasons PZ writes about. But it has generated a good deal of “ado”. The HGP has given us an enormous body of evidence for a great many other questions about the genome; its structure, its correlates to disease and phenotypes, genetic relationships inter and intra species, answering (and more importantly generating) questions about evolution, etc. Maybe I’m not getting your objection to HGP clear.
No, actually, the HGP was vastly over-hyped from the very beginning. There was lots of talk about reconstructing humans from genetic information, deciphering what makes us human, etc.
But I agree it wasn’t nothing. It’s actually a very useful tool. It’s just not the answer.
Ooh, I love this kinda stuff, PZ!
One question comes to my mind: Did having meatier fins make for stronger swimmers? And if so, why are there still so many fish with puny rayed fins? Was it maybe a trade-off between stronger swimming versus maneuverability?
Reminds me a bit of my comments in this thread: http://freethoughtblogs.com/pharyngula/2013/02/03/another-really-stupid-argument-from-william-lane-craig/
The same craving for a single immediate cause leads us to the blueprint metaphor and to infer proximal, mental causes for complex behaviors extended in time.
Good stuff.
PZ wrote; “here was lots of talk about reconstructing humans from genetic information, deciphering what makes us human, etc.”
OK, I have to admit, back when the HGP was being promoted I was buried in T cell differentiation. I don’t think I looked up from mouse thymuses for like five years. So I probably missed all that.
There are some very specific gene clusters that do follow the “blueprint” model (e.g. the Hox genes), but they are (1) a minority of the genome, and (2) less blueprint-y than frequently believed. Yes, disruption of the order of Hox genes does lead to abnormalities (Antennapedia being the classic example), but this is because of the specific function of the Hox genes – they lay out the central anterior-posterior axis and map out where specific clusters of cells go. Once those cells go there (as directed by the Hox patterning), they then do their job – and thus you have legs growing where the antennae should be, because the transposition has led to the laying-down of a “leg” signal where the “antenna” signal should be. The leg construction is based purely on the cells following the cues of their local environment, not them going back and reading some central “blueprint” of the body.
Between “blueprint” and “recipe,” I’d pick recipe as a better metaphor for evolution. Recipes tend to have more wiggle room than things built with blueprints, so it’s probably better for cases where you’re trying to drill in the idea that variation is natural, whereas blueprint tends to imply a finely honed machine that might fall apart if you take out or change key parts. An engineering design failure can easily be catastrophic to the system, whereas a culinary failure is more likely to result in something less appetizing while still edible. You can think of development as a chef (mindlessly following the recipe he was given), the produced organism as a meal, and natural selection and such as the judges of the cooking contest.
But, since we’re specifically talking about development of an organism, the chef is a big part of what we’re trying to explain, so it leads into something like the homunculus fallacy in psychology. You can’t explain what’s going inside a person’s head by likening various mental processes to a little person inside his head. “Person” is the phenomena to be explained, not an explanation.
A colony of multi-task robots is probably a better metaphor for development. I think an insect colony might also be workable. They aren’t centrally controlled. Ants respond to the environment, send signals to other ants who adjust their behavior, which adds up to complex colony behavior. Different environment, different responses. You can affect an ant colony’s behavior by spraying their communication scents in the right places. In this case, I presume those scent communications are equivalent to triggers activating or suppressing relevant gene expressions, changing the cell’s behavior.
Random aside: One phenomenon I’ve heard of with army ants is called a “circular mill” where a colony ends up in a loop, running in circles until they die of exhaustion because each ant is simply following the one in front of it on the assumption that there’s ants in the lead, making decisions based on the environment.
Not really, because ray-finned fish get their swim power from the side-to-side motion of their meaty bodies and tails, while the fins are mainly for fine maneuvering. Things that do swim with their ‘fins’ look like rays and skates, with big fleshy wings. (Or penguins, which swim with paddles that are actual wings, powered by wing- in ancestors, leg – muscles.)
I’ve always hated the blueprint analogy too. It just leads to confusion, and creationists and IDers bang hard on that to make it look like evolution is impossible.
I just started reading Your Inner Fish (thanks to the people who have talked about it here) and, in fact, was looking at one of those “compare the forelimb/fin structures” drawings just yesterday. For me, a nice coincidence.
An important thing to remember is that zebrafish do not retain the ancestral condition. They’re teleosts, and as such they have lost the distal part of the fin skeleton. This is why such work is now occasionally done on paddlefish and sturgeons and small sharks. Zebrafish are model organisms, they’re easy to breed in a lab, their genome is very well known, but they’ve gone their own way, and they’ve gone it quite far.
The drawing of the fin of Panderichthys has been outdated for over four years now. The fossil is poorly preserved and prepared, so it had been misinterpreted. It was CT-scanned and published in 2008. The terminal “plate” of the endoskeleton actually consists of several bones, making the whole thing look a lot more like Sauripterus or Tiktaalik.
Would surprise me.
Could be.
Not even. It’s a template for a set of things that have very complex electrostatic attractions to and repulsions from each other and catalyze a variety of chemical reactions.
The fin endoskeleton, I mean.
Lovely, informative post, PZ. Straining for analogies (blueprint and recipe the genome is definitely NOT) is a constant problem. Bits of the way these things work resembles execution of a computer program in multiple threads, but isn’t the closest analogy we have something like the local rules that, e.g., birds follow in flocks and which give the whole flock its amazing morphology (both shape and location changes). Each bird is responding to a small set of rules depending on its immediate neighbour’s behaviour and other clues.
I’m also reminded of Douglas Hofstadter’s brilliant dialogue involving an anteater. “Why do you eat the ants?” Because they spell out “Eat me” in anteater script. Hofstadter was talking about levels of computer program, with machine code at the lwoest level. If the genome is the lowest level machine code program, then the factors that interact with promoters, epigenetic inputs and so on behave like higher-level inputs, all determining an outcome that’s not predictable by reading the machine code.
I wonder it would be easier to explain to people if you use plants as a model (or even metaphor) instead of vertebrates? Vertebrates are what most people are interested in, of course, but quite a lot of people are familiar with the concept of intervening with young plant growth and observing the effects on the mature plant.
In racking my brains for a better metaphor than “blueprint” or “recipe,” plant growth came to mind. Of course, that’s pretty much the very same problem that we are trying to explain anyway. But lots of people mess with plants and it is easy to do at home in ways that animal studies are not.
nooneinparticular said it in #3, Posts like this why I keep coming back to Pharyngula. Same here. Thanks, PZ, one keeps learning here.
WRT metaphors, my interpretation of cellular automata is hardware, programmed by some other hardware (the genome) that is written in some BASIC, with a lot of GOTO, IF…THEN…, GOSUB, and above all, PEEK and POKE commands. And yes, it is possible to have recursion* in BASIC, as long as you build the proper stack.
_____________________
In my youth I programmed a 64-disc Tower of Hanoi on a 48K dedicated-BASIC machine and I could prove it wouldn’t crash until all 64 discs were at their destination. I’m still proud of that, since my Pascal teacher told me that it wasn’t possible. That got me going.
That sounds good to me.
I know this isn’t the point of PZ’s posting, but the bit about stuffing the pita and future evolutionary pressures would shape it into something useful seems like a just so story. Not that I doubt for a second that evolution happens, but changes which are costly or harmful without benefit would very likely result in a dead end.
Is there any speculation on why filling the pita would be advantageous that future revisions of coherent bone and musculature make it obviously useful?
Thanks for this very, very interesting post, Professor M!
This is just another reminder of how beautiful and amazing the process of evolutionary biology actually is.
So much richer than “The Sky Fairy did it.”
Thanks for this.
Tetrapods descended from the so-called lobe-fins, which today include lungfish and coelacanths. This fish always had “meaty” fins. The teleosts are the most successful modern clade of the other great fish family, the so-called ray-fins.
There are I think 3 species of surviving lungfish and 2 species of coelacanths.
There are more teleost species than all other vertebrates combined.
In the evolutionary battle of swimmers, it would appear the ray fins kicked the lobe fins’ butts, and, to indulge in a little teleological metaphor, chased the bedraggled lobe fin survivors right out of the water (though whales, mosasaurs, pliosaurs, sea turtles, penguins, seals, ichthyosaurs, and Michael Phelps all count as lobe fin swimmers).
Though of course, fin anatomy on its own is not likely to be the only reason for the teleosts’ current relative success.
And so, pertaining to this, it is my understanding that ray-fins (Actinopterygii, and they include more than just the teleosts) did not evolve from lobe-fins (Sarcopterygii), but that the two clades are sister-clades sharing a (as yet unknown) common ancestor. The question would be what was the fin-type of that ancestor, and was Hoxd13 expression high or low in the limb in it? So would it have been the addition of enhancers to the ancestral state that produced what went on to become the tetrapod limb, or was it the removal of enhancers from the ancestral state that went on to produce the teleost ray-fin?
Of course if you go back far enough, sooner or later you reach an ancestor with no limbs at all, so you must have started at some point from a state where there would have been no Hoxd13 expression at all.
But given what we already know about how gene networks tend to evolve, it seems to me to most likely hypothesis to start with is that the very first fins began with a general expression of a whole bunch of closely related duplicated Hox genes with overlapping and comparatively poorly defined functions, followed by specialization and paring down of these duplicated genes. ie, the expression of the original ancestral gene of Hoxd13 probably started out high-ish in the primordial limb/fin.
@David Marjanović #25:
Well, yes: GIGO.
On a complete aside, earlier today I took a look at the web site of a company who might, in the near future, be supplying software to the organisation I work for. Their marketeers had gone beyond the stock, empty phrases such as ‘customer service is in our DNA’ to listing what was ‘in their genome’.
After the wave of nausea had passed I wondered if they’d end up describing their software as their phenotype. But I don’t think I want to find out.
Damn my lazy lack of previewing! That should read ‘in our genome’.
It is only if you take the teleological metaphor within the description literally.
Consider the ancestral condition – a finless worm-like form. On the flank the anatomy will consist of a layer of ectoderm with a layer of mesoderm below it. For a limb/fin to form, what has to happen is for a part of that outer layer to fold out.
Since having any sort of fin is in the right circumstance an advantage over no fin at all, the actual folding mechanism that produces that first proto-limb/fin/appendage does not have to be all that precise. Since it takes a certain degree of precision to guarantee that only ectoderm folds and mesoderm does not, the most likely scenario is that the first primordial fold included both ectoderm and mesoderm within it, perhaps haphazardly.
Thus the starting condition is a pita that is partly filled. And from that starting condition we’ve actually seen both alternate processes occurred. In some lineages the pita has been filled some more. In others the pita has in fact been emptied. And both are successful in their own different ways.
But to see where refinements of the interior of the pita might be useful one simply has to realize that a proto-limb/fin that can move somewhat is superior to one that can not. The primordial mesoderm would have included muscle tissue, since muscle is one of the things that mesoderm does. Thus the presence of mesoderm in the original pita would have provided some mobility. Thus even an incoherent lump of mesoderm that can do nothing more than contract and quiver a bit is already potentially useful in the right circumstances, and from that starting point selection can get started.
ok, so i have used this analogy in my secondary biology classroom. Referring to DNA as the bio-chemical blue print of our bodies that can be “read” and “constructed” through the process of transcription and translation. I mention that all of my analogies are imperfect, but is this one particularly grievous, is this doing more harm than help? And does anyone have a better “fitting” analogy? thanks for any responses :)
A gene could be said to be a blue print for a protein and/or RNA transcript (though tertiary and quaternary structure still throws a wrench into that metaphor), but a genome is not a blueprint for an organism.
And only a tiny (in eukaryotes, less so in prokaryotes) proportion of any genome consists of genes.
I can see the problem with your explanation right here: it takes you 185 words to stress the importance of DNA. On the other side, someone comes and say “hey DNA is like a blueprint”: shorter sentence wins.
Why not treating the DNA=Blueprint like the Big Bang? The universe did not start in a big Hollywoodian explosion, but it’s not a bad way to allow neophytes to picture it and realize that it’s a big deal.
Blueprint? Recipe? Map? Why not something less archaic, simpler, closer to everyday life. Why not “algorithm”?
Grasping for analogies myself. Listing all the elementary particles and their interactions is not a “blueprint” for, say, aerodynamics. Paradigm hierarchy?
runrabbitrun, amphiox
Metaphors are by their nature models and necessarily inaccurate. They are only useful in they relate an unfamiliar concept to a familiar one. In that much of the information required to specify developmental pathways is encoded in the DNA, I think likening DNA to a blueprint for building an organism can be useful if this concept is all that you wish to convey*. At an intermediate level, I think it may be more appropriate to say that the genome is a blueprint for building proteins, among other things. And so on. As long as the metaphor is conveying whatever aspect of genome function that you are trying to explain, it may be useful.
I teach at a college level, and at least in instances like this, I just abandon metaphor whenever possible (because these metaphors becomes too complex to be that useful). My difficulty is more in deciding which generalizations are useful.
*The kind of discussion you may have with an elementary school student, I guess.
“DNA is like pie.”
I win!
“If you want to make an apple pie from scratch, you must first create the universe.”
Since I first heard of epigenetics, I’ve analogized DNA to a huge pad of checklists. The intergenerational portion of epi is list items that come already checked off, but the genomic part (the majority) is clean.
Each cell division issues about half the pad to each successor; this half has some epigenetic mods checked off — these are the items on the list that the other half is to take care of. Sometimes pages get messed up, or even go missing — these events can cause developmental problems.
I’m sure that someone who knows his stuff could make a decent metaphor out of this, should they feel it worthwhile.
I’d still like PZ’s insights as to my PEEK and POKE comments @24.
Here’s what I tell my students – DNA is not a “blueprint”, it’s a “parts list”. A blueprint suggests there is a plan, a set of instructions, and if the builder follows these, he or she will end up with a house. But DNA is really just giving us the parts (the proteins) – we’ll need shingles, siding, nails, copper wire, PVC pipe, cement, glass, etc. Now we have the picture of a bunch of parts getting dumped into a big hole at the construction site, and you come back 9 months later and – holy cow! A house!
A flawed analogy too, but one that hopefully opens the students’ minds to thinking about complex interactions between cells and between organism and environment during development.
Nah, it’s not a programming language, either.
I’m guessing that people have a hard time with the idea that the cells with their DNA in the body are really behaving like automata like an independent organism. There is more of a communal nature to what is happening is it not? A highly integrated colonial organism.
I have always had a compulsion to come up with a “better” more a accurate description, metaphor simile what ever and what you were describing sounds some what similar to what in Jazz is called a Head Arrangement everyone knows more or less how the tune goes and how their sections are arranged their parts but each has some leeway to their interpretation and their reaction to what the rest of the band players are doing often in a jam session some very novel improvisations can develop depending on the players involved.
I like how in biology as in much of all science the edges seem to bluer and the detail refinement just keeps going on and on and the simple if not still primitive way we make things and like to see things as discrete entities is not particularly accutate.
Thank you for more clearly show us that we are just scratching the surface and we ain’t found “the answer” yet
uncle frogy
From the announcement for a talk I wish I could be going to, but is on the opposite side of the planet from me:
Sweet science lesson! I’ll go to bed noticeably smarter tonight than I did last night. Seems like I have fewer and fewer days when I can say that lately, so I appreciate the knowledge boost.
Damn, my lunch hour is almost over; I shall have to reread and study this blog post later this evening. Did anyone else see that bright flash of light? That was me understanding how, as a result of the first several paragraphs, that DNA is not a “program” for some sort of Turing machine. If anything, it’s the most horrid spaghetti code, devilishly difficult to debug and next to impossible to fix: any fix you make will break something else. (That’s why all those drug commercials have long, scary lists of side effects.) DNA code is not nicely compartmentalized into modules. It’s not a blueprint; it’s not a program.
PZMyers, I think that this popular misunderstanding of how DNA functions is the single biggest impediment to the common understanding of biology and evolution. I think you have the subject of a book here, one with the same potential effect on general understanding of biology as Dawkins’ [i]Selfish Gene[/i].
At the very least, it’s the subject of an interesting panel in the science track at a science fiction convention!
If this is referring to the metaphor of computer code, it was never ever any good. Sections of DNS are jigs/templates, and the rest is mostly quite a lot of spacer between these tools on the rack.
kevinkirkpatrick @ 10
Not really.
Think about classes, each with their own attributes and methods, that interact together without central oversight.
«WRT metaphors, my interpretation of cellular automata is hardware, programmed by some other hardware (the genome) that is written in some BASIC, with a lot of GOTO, IF…THEN…, GOSUB, and above all, PEEK and POKE commands. And yes, it is possible to have recursion* in BASIC, as long as you build the proper stack.»
The essential elements of the cellular automaton metaphor are that you have a bunch of independent units which are state machines, which are all running the same program and which are spatially linked to each other. The state of each individual unit is determined at any point in time by the way its programing, given its current state, interacts with the environment (in the case of cellular automata “environment” means the state of other cells –usually immediate neighbors– but in the case of biological systems “environment” has a broader meaning). The individual units are not going to behave identically at any given time because, at any given time, they are going to be in different states (due both to being exposed to different environments and to having a different state history) and because they are going to be in a different environment. There will be many emergent behaviors but they are not in any way explicitly programmed (this is a key point, in my opinion –thus my clever use of the bold HTML markup and my use of repetition for emphasis).
This is why, in my opinion, the ‘genome as computer program’ metaphor falls way short and the cellular automaton metaphor being used by P. Z. Myers works much better. The issue is not that the biological system is not a Turing machine (though it is not a Turing machine) but rather a set of interlinked Turing machines (though it is also not a set of interlinked Turing machines –which would actually be computationally equivalent to a Turing machine, in any case). The issue is that when one uses the ‘genome as computer code’ metaphor it tends to imply a direct encoding of the developmental patterns which does not really exist in the biology itself and which tends to neglect the environmental aspects with respect to the individual cells of the developing system and with respect to the system as a whole.
The prototypical cellular automaton system is John Conway’s game of life (see http://en.wikipedia.org/wiki/Conway%27s_Game_of_Life ). As I understand it (not having read it), Stephen Wolfram has written a book basically purporting to use cellular automatas to answer the great question of life, the universe and everything.
I have a question which might fall into some kind of lmgtfy/rtfm/read the FAQ/etc. category. If it does, my apologies.
Are there any viable software projects out there (preferably of the FOSS variety) designed to simulate aspects of biological development? If nothing else it might be fun to create a “Cambrian explosion” in silico by tinkering with such a system. At best, you might be able to simulate aspects of some real biological developmental systems and even use such a software to generate new, plausible hypotheses to direct real laboratory research using molecular biology tools (like the research reviewed in this blog post).
I can think of two general approaches one might use to create such a software system (although I have no idea if either or both would work): one approach would be to use a strict cellular automaton system such that each cell in a 3-D lattice represents components of much larger pieces of the real biological system it is meant to simulate (so a computational cell might have extracellular matrix states, cell membrane states or cytoplasmic states –that is, an organismal cell would actually be composed of a largeish number of computational lattice cells) and the other approach would make each individual organismal cell be represented by a an individual computational cell but the system would not be a true cellular automaton* in that individual cells would be able to vary their spatial characteristics and their spatial relationships to each other (just as is the case with the real biological system). The physics of growth, adhesion, signal gradients, etc. would have to be encoded into the programming of individual computational cells along with other aspects of the system.
* A problem I could foresee with this more “literal” approach is that it might be difficult to model a cytoplasmic determinant as being located in one end of a cell.
The analogy my Developmental Biology professor used is that development is like ecological succession. There’s obviously no master plan, but the various communities of organisms interact with each other and the environment to produce a dynamic system. Eventually you reach some end stage (a climax forest or an adult animal) that looks much the same as any other. There will be slight differences depending on the environment, what organisms colonized the place in what order, and a fair bit of chaotic noise.
This obviously doesn’t work if someone is unfamiliar with ecological succession. I found it really useful, and hopefully some of you will too.
I am personally a fan of the recipe analogy. But I think the more defining point is that genes aren’t working in a vacuum or through a linear progression of steps–they participate in a massively simultaneous set of processes.
You just replaced a metaphor you don’t like with a non-sequitur: this veers very close to a chewbacca defense
“There will be many emergent behaviors but they are not in any way explicitly programmed”
I’m failing to see that distinction. You mean they don’t have a programmer? That’s obvious. But otherwise it is just as easy to create computer code where individual elements behave in a similar way. As in the “game of life” problem (a cellular automaton simulation with emergent behaviours arising despite there being only a simple set of rules).
Code that behaves like this is actually used in speech, text and image recognition. To me the analogy falls short not because computer code can’t do the kind of things that DNA does, but because you don’t usually get that kind of computer code producing behaviour as complex as a human cell dows and it would be grossly inefficient to write that kind of code for the majority of programming tasks. “Typical” code produces results nothing like DNA does, and in completely different ways.
That was interesting! Thank you for explaining it clearly enough that a computer techie understands.
3 genera – but there are 4 species of the African genus (Protopterus), or 7 if you want to elevate the subspecies recognized on Wikipedia to species rank.
And things like the particularly mobile jaw apparatus of rayfins don’t look like the whole story either.
In particular, the mass extinction at the Devonian/Carboniferous boundary eliminated all lobefins except for coelacanths, lungfish, rhizodontids, megalichthyids, and tetrapods (in a very wide sense); the rhizodontids and most coelacanths disappeared sometime close to the end of the Carboniferous, and the megalichthyids at the end of the Early Permian or so. In the Devonian, the diversity was huge.
The paddlefish and sturgeon I mentioned are actinopterygians but not teleosts; they have a lot more pita filling than a zebrafish does. And the bichirs, the sister-group to all other extant actinopterygians… I recently asked a creationist what happens when you take one of those out of the water: it walks. Look at these forefins.
Am I allowed to say that DNA is an instruction set?
You know, kind of like “Do this little thing if this thing comes around and do that little thing if that thing comes around.” Like a computer program trying to do complex things off of loops of simple rules. And then when they’re all combined they do all sorts of weird combinations of things depending on the surrounding environment and poof! – it’s a baby.
Is that a better layman description? Because that’s the one I use.
Pretty much the latter is the case. It’s a given that biological cells don’t have a programmer. So you are suggesting that P. Z. Myers is saying that DNA is not like computer program but that, instead, it’s like a specific kind of computer program (since, indeed, cellular automata are a class of computer programs)?
You are correct but you are also being a little bit pedantic and possibly missing the point. First of all, this is meant as an analogy and not as a suggestion that one thing is exactly like the other. Also, the sort of AI approaches you mention (things like genetic algorithms, neural networks, various other learning methods, etc.) are absolutely not what a layperson thinks of when they think of a computer program. The real point, though, is that, for example, in Conway’s Game of Life there exists no rules on how to create a glider and yet, under certain circumstances, gliders emerge in its “developmental processes”. Likewise, there exists no sequences dictating the rules on how to make an arm in your DNA.
In a cellular automaton you have a set of rules built into each cell. These rules do not even make sense until you insert the cell into the context of an n-dimensional lattice. Likewise, in a biological cell you have a bunch of instructions on how to make proteins. On their own (that is, as a raw sequence of DNA) they don’t even make any sense (if you simply made all the peptides & RNA’s which are encoded in the sequence you would not get anything useful). Only when the whole cell (and the cell itself is also part of the context of DNA) is inserted into its proper biological context can the emergent properties of developmental processes within that particular context be actualized.
I love these posts, PZ: it’s great to have the opportunity to learn more about this.
He he.. Programmers hell:
10 if blah then do stuff : goto 30
20 if blah2 then do stuff : goto 500
30 if blah3 then do stuff : goto 10
40 …
Nearly impossible to debug, and… usually, beyond the first time any competent teacher said, “You have the option to use goto, but just.. don’t. There are some very specific cases where its faster, and more efficient, but, usually, it just makes things more complicated, such as by created a condition where the code will ‘never’ exit.” Sort of what happens when “copy” gets turned on in a gene, and the “end” statement is missing (the equivalent, apparently, of telling genes, “keep processing this code, until I *never* tell you to stop.”
Ur.. meant to end that first long sentence there with, “gets you an F.”, if you like, use it, instead of better logic. lol
I was disappointed to see this quote hadn’t already been mentioned:
“Remember, genes are NOT blueprints. This means you can’t, for example, insert “the genes for an elephant’s trunk” into a giraffe and get a giraffe with a trunk. There are no genes for trunks. What you can do with genes is chemistry, since DNA codes for chemicals.”
— Academician Prokhor Zakharov, Sid Meiers Alpha Centauri
It’s remarkable to me that a 14-year-old video game does a better job of explaining science than institutions like the NHGRI today.
The Evolution concept in Turkish-Islamic people
http://www.sonsuzlukkulesi.com/evolution_turk_muslim/