Now you’ve got another paper you can file with that dead salmon fMRI paper: one that analyzes the transcriptome, or excuse me, the thanatotranscriptome, of dead zebrafish and mice.
You should not be surprised to learn that when a multicellular organism dies, it’s not as if every single cell is abruptly extinguished: the integrated, functional activity of the individual as a whole ceases, but individual cells struggle on for a while — they’re not getting oxygen or nutrients, in a mouse they’re experiencing thermal stress as the body rapidly cools, but it takes a while for all of the cells to starve or suffocate or undergo necrosis. It seems to take a couple of days, actually. You can measure the declining amounts of RNA present in the dead animals, and yep, it looks like everything is done after a few days. This kind of study has also been done in human corpses, which show that RNA transcription continues for a couple of days.
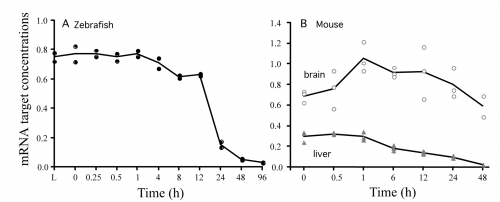
Total mRNA abundance (arbitrary units, a.u.) by postmortem time determined using all calibrated microarray probes. A, extracted from whole zebrafish; B, extracted from brain and liver tissues of whole mice. Each datum point represents the mRNA from two organisms in the zebrafish and a single organism in the mouse.
This paper went a bit deeper, though, and asked what genes are being transcribed in the dying cells of the dead mouse and fish. Some genes are actually actively upregulated, so it’s not as if they all just stop and decay on the instant. And, well, it’s a bit unsurprising which ones are switched on: genes involved in stress responses, metabolic activity in reaction to hypoxia, genes involved in inflammation, and immune system genes. Starved cells are stressed, and clearly decaying bodies are being invaded and exploited by bacteria, so the immune system is triggered. Inflammation is a response to injury, and I guess we’d have to say that death is a rather serious injury.
Another activity we take for granted is cell-level transport: your cells are very busy trying to maintain a constant osmotic environment. In death, cells are suddenly leaking, and the extracellular salt concentrations are changing, so yes, again unsurprisingly, dying cells are making a futile struggle to maintain their salt balance, and are desperately switching on all kinds of pumps.
The authors did not expect various developmental genes to be activated by death, but I’m not at all surprised. The inactivation of specific genes is also an active process — many of these switches are not at all passive, but require ongoing regulatory activity in cells to maintain their state. So at death a whole bizarre, seemingly random collection of developmental genes are increasingly transcribed. What I suspect we’re seeing is a gradual release of active deregulation — that some aspects of the normally active epistatic circuitry are being spuriously switched on as other control elements are falling apart. It’s like when the various controls on the bridge of the Enterprise start sparking and catching fire when the Klingons hit them with a phaser burst — there’s probably a pattern to it that reflects the underlying wiring.
The authors argue that there’s some utility to this study. It might be a useful assay for forensic research — which genes are upregulated could tell you something about the time of death, for instance. They also make a case that this is useful information for resuscitation and transplant research.
We initially thought that sudden death of a vertebrate would be analogous to a car driving down a highway and running out of gas. For a short time, engine pistons will move up and down and spark plugs will spark — but eventually the car will grind to a halt and “die”. Yet, in our study we find hundreds of genes are upregulated many hours postmortem, with some (e.g., Kcnv2, Pafr, Degs2, Ogfod1, Ppp2rla, Ror1, and Iftm1) upregulated days after organismal death. This finding is surprising because in our car analogy, one would not expect window wipers to suddenly turn on and the horn to honk several days after running out of gas.
Since the postmortem upregulation of genes occurred in both the zebrafish and the mouse in our study, it is reasonable to suggest that other multicellular eukaryotes will display a similar phenomenon. What does this phenomenon mean in the context of organismal life? We conjecture that the highly ordered structure of an organism – evolved and refined through natural selection and self-organizing processes – undergoes a thermodynamically driven process of spontaneous disintegration through complex pathways, which apparently involve the upregulation of genes and feedback loops. While evolution played a role in pre-patterning of these pathways, it does not play any role in its disintegration fate. One could argue that some of these pathways have evolved to favor healing or “resuscitation” after severe injury. For example, the upregulation of inflammation response genes indicate that a signal of infection or injury is sensed by the still alive cells after death of the body. Alternatively, the upregulation may be due to fast decay of some repressors of genes or whole pathways (see below). Hence, it will be of interest to study this in more detail, since this could, for example, provide insights into how to better preserve organs retrieved for transplantation.
Eh, I don’t find their surprising result at all surprising, but OK, this could be useful information. I don’t think this is a very good way to study evolution or development, though — as they note, the activity of these genes is not a consequence of their selection, but of their functional role in life, and their activities in such a literally pathological state as death are not going to reflect how they were shaped in their formation.
By the way, if you’re concerned about the grisly details of how these animals were killed (and you should be!), the mice were executed by snapping their necks, which is very, very quick, and the fish were killed by cooling them (which is the recommended method for humane termination among aquarists) and then dropping them in liquid nitrogen.
Alexander E Pozhitkov, Rafik Neme, Tomislav Domazet-Loso, Brian Leroux, Shivani Soni, Diethard Tautz, Peter Anthony Noble (2016) Thanatotranscriptome: genes actively expressed after organismal death. bioRxiv doi: http://dx.doi.org/10.1101/058305.


Do zebrafish have souls?
Don’t Klingons use disruptors instead of phasers? ;-P
So, in the spirit of most science journalism, the obvious thing to do here is to read only the headline, and jump to the conclusion that… zombies are scientifically confirmed to exist!!11!!Eleventy!
Yes. It’s a large, complex neuron located at the hindbrain/midbrain boundary.
Didn’t everyone know that?
Well, because no one else has jumped on this yet, I’ll have to add the obvious movie reference:
“There’s a big difference between mostly dead and all dead…Now, mostly dead is slightly alive. Now all dead, well with all dead there’s usually only one thing you can do…Go through his clothes and look for loose change.” Miracle Max
https://youtu.be/xbE8E1ez97M
This really is a fascinating paper and one I’ll be using parts of for my honors biology students come the fall when we’re discussing cell functions and multi-cellular life. Thanks for posting this PZ.
Just, on a tangent, I object to their analogy:
“This finding is surprising because in our car analogy, one would not expect window wipers to suddenly turn on and the horn to honk several days after running out of gas.”
What does wipers and the horn have to with it running out of fuel? If those functions weren’t working after several days, the problem would be the battery.
Trying to think of a different analogy, but my imagination fail me.
At least we can now quantify why Dr. Herbert West’s subjects just “weren’t fresh enough”.
I could never make sense of the paper, so riddle me this you folks who know: In the chart for the zebra fish and the mouse liver there’s a steady decline the mRNA abundances, but for the mouse brain tissue it appears there’s an increase with a spike after about an hour before a steady decline. Just curious what that indicates.
I was at a lecture from an itinerant English professor from GB. Her area was bio-semiotics. There were a lot of very questionable arguments being presented, but I tried to keep an open mind as her express goal was to eliminate some of the stigma and hostility humanities folks have for the hard sciences.
The main argument was that the Human Genome Project proved that DNA and genes were not sufficient to explain how organisms come into being. The “neo-Darwinist” blueprint model is inadequate. It’s much more complicated. Leading biologists (interviewed in a book she held up titled something like the “New Paradigm”) have all concluded that the “Neo Darwinists” were wrong, but that there is a conspiracy in academia where folks like Richard Dawkins are keeping the new paradigm folks from getting published.
What was really going on was something like sign interpretation in cells (the semiotics), and that because information is not material, exclusively material explanations for inheritance and the development of organisms has failed, despite the dogmatism of the Neo-Darwinists.
The conspiracy claim was enough to raise my eyebrows, and history has not been kind to arguments of the form, “this is too complicated for current explanations, ergo it must be some immaterial something or other”, but I did realize that I have a very poor understanding of the current science on the subject. What would you folks recommend on the subject? Is this an argument you’ve encountered before?
Just another “god of the gaps” argument. God is very small these days, and the gaps are getting smaller.
mandarb, #6,
What bugs me the most about that analogy is that genes are not being upregulated “several days after” the cells run out of fuel. Cells’ fuel is ATP and I’m pretty sure there’s no transcription going on after the ATP pool becomes thoroughly depleted. Also, window wipers turning on and the horn honking are not mechanisms used by cars to cope with running out of fuel, while the genes being turned on here are mostly related to mechanisms used by cells to cope with decay-related stress.
So, maybe:
Except this would not be surprising at all!
Still, it’s a very interesting study.
@ Nerd of Redhead, Dances OM Trolls
Agree completely.
robro @8
It is the soul packing it’s stuff and leaving the body.
Actually, I would expect that the RNA activity post-mortem is the result of selection…just not selected for by death. I’m sure there’s some very interesting work to be done on which genes and processes get activated and it will be strongly related to evolved cell stress responses.
doubtthat@9:
OT but my response is that the person speaking is one of those who uses their own area of expertise to pontificate on other areas that they don’t understand at all. There’s nothing wrong with using semiotics as an approach to transcription because there is a lot of message-sharing going on, so I’m sure there are some interesting semiotic models that could be relevant. BUT, she doesn’t seem to know that information is materially encoded and is not magic, or that we understood that DNA and genes were not sufficient to explain development and embryology long before the Human Genome Project. Really, this is just the touchy-feely flip side to neo-liberal evolutionary psychologists who use their own (often sketchy) understanding of evolution to stomp all over anthropology, sociology, economics and so on.
Interesting post. I always learn something new when I visit Pharyngula.
By the way, PZ, I enjoyed reading your article from a biologist’s perspective in the “Death Issue” of Free Inquiry.
Re: Robro (#8):
I only have the chart from PZ’s post to go by, but the ‘spike’ doesn’t look like that much of a spike to me. Unlike the other data, the mouse brain data points seem to be spread out a fair bit. Only the point immediately after the peak has three closely spaced data points. The ones from the peak and the point before deviate a lot. It’s not too difficult to imagine the graph might look differently (lower peak, point before it slightly higher?) if you repeated the experiment to get a new triplicate measurement.
On the whole I’d expect a curve (shape) very similar to the mouse liver data, where the interesting take-away (as a non-biologist) would be the higher abundance. But then this might be perfectly normal when comparing brain and liver in any organism, late or otherwise.
i love how the first two comments captured the full ontological spectrum of grand, unknowable truths, and minute inconsistencies with star trek canon.
@chrislawson
There was definitely a combination of potentially interesting insight and stuff dangerously approaching the worst kind of woo.
As I said, it did make me feel like my substantive knowledge of the topic was woefully out of date. Are there good layman’s works on the subject? Bryson-style? I could use a good overview on the current understanding of cell biology and genetic.