Casey Luskin is such a great gift to the scientific community. The public spokesman for the Discovery Institute has a law degree and a Masters degree (in Science! Earth Science, that is) and thinks he is qualified to analyze papers in genetics and molecular biology, fields in which he hasn’t the slightest smattering of background, and he keeps falling flat on his face. It’s hilarious! The Discovery Institute is so hard up for competent talent, though, that they keep letting him make a spectacle of his ignorance.
I really, really hope Luskin lives a long time and keeps his job as a frontman for Intelligent Design creationism. He just makes me so happy.
His latest tirade is inspired by the New York Times, which ran an article on highlights from the coelacanth genome. Luskin doesn’t think very deeply, so he keeps making these arguments that he thinks are terribly damaging to evolution because he doesn’t comprehend the significance of what he’s saying. For instance, he sneers at the fact that we keep finding conserved elements in the genome, because as we all know, there are lots of conserved elements.
Hox genes are known to be widely conserved among vertebrates, so the fact that homology was found between Hox-gene-associated DNA across these organisms isn’t very surprising.
Stop, Casey, and think. Here’s this fascinating observation, that we keep finding conserved genes and conserved regulatory regions between mice and fish, which ought to tell you something, and your argument against a specific example is that it isn’t rare? It really tells you something when your critics’ rebuttal to a piece of evidence is that you’ve got so much evidence for your position that they’re tuning out whenever you talk about the detais.
This is Luskin’s approach to every example given in the NY Times article: ‘Yeah? So? There are homologous genes all over the place!’ I think Luskin might just live forever, which thrills me to pieces. He could get into a running gun battle with a mob from Answers in Genesis, be riddled with bullets, and he’ll just point to a nick in his ear and say, “Yeah, so? This one didn’t kill me!” and then dismiss all the other wounds because they’re so common that no one should care any more. It is truly the logic of immortals.
The specific example he’s addressing in his dismissal of Hox conservation, though, is a region of DNA that may play a role in mammals in the formation of the placenta. Luskin pooh-poohs the relevance of this observation by highlighting what we don’t know, rather than the evidence at hand.
The authors aren’t sure exactly what this particular segment of DNA does, though it’s probably a promoter region. In mice the corresponding homologous region is associated with Hox genes that are important for forming the placenta. Ergo, we’ve solved the mystery of how the placenta evolved. Right?
Not really. Again, all that was found was a little homologous promoter region in Hox-gene related DNA in these two types of organisms. Given that we don’t even understand exactly what these genes do or how they work, obviously the study offered no discussion of what mutations might have provided an evolutionary advantage. No evolutionary pathway was proposed, or even discussed. So there’s not much meat to this story, other than a nice little region of homology between two shared, functional pieces of Hox-gene-related DNA. But of course, such shared functional DNA could be the result of common design and need not indicate common descent or Darwinian evolution.
He’s right that this genetic sequence does not “solve the mystery” of the placenta, but then the authors of the original sequence analysis paper do not claim that it does. There’s a lot that we don’t know about the translation of any genetic sequence into a morphological feature, but that doesn’t mean we know nothing — the paper is talking about stuff that we actually learned from comparing the genomes of different organisms.
We have identified a region of the coelacanth HOX-A cluster that may have been involved in the evolution of extra-embryonic structures in tetrapods, including the eutherian placenta. Global alignment of the coelacanth Hoxa14–Hoxa13 region with the homologous regions of the horn shark, chicken, human and mouse revealed a CNE just upstream of the coelacanth Hoxa14 gene. This conserved stretch is not found in teleost fishes but is highly conserved among horn shark, chicken, human and mouse despite the fact that the chicken, human and mouse have no Hoxa14 orthologues, and that the horn shark Hoxa14 gene has become a pseudogene. This CNE, HA14E1, corresponds to the proximal promoter-enhancer region of the Hoxa14 gene in Latimeria. HA14E1 is more than 99% identical between mouse, human and all other sequenced mammals, and would therefore be considered to be an ultra-conserved element. The high level of conservation suggests that this element, which already possessed promoter activity, may have been coopted for other functions despite the loss of the Hoxa14 gene in amniotes
What they are saying is that they found a region of DNA that is remarkably highly conserved between coelacanths and humans, and that among them are Conserved Non-coding Elements (CNEs), DNA that is not translated into proteins but instead is likely to be important in switching genes off and on. What they found is that this switch is linked to a particular gene, Hoxa14, in the coelacanth which is completely absent in mammals…but the switch still exists, but is now coupled to a gene associated with the formation of extra-embryonic membranes, like the placenta.
The key point is the known information: this snippet of DNA is highly conserved across all vertebrates, but the gene it is associated with has decayed in sharks and disappeared entirely in tetrapods. The question is how the switch has persisted: Luskin’s preferred explanation is that God spliced this particular piece of DNA into every vertebrate species; the scientific explanation is that it shows a pattern of shared history, and that its role is conserved in tetrapods because it has found a novel function in regulating a different gene.
So far, this is nothing but Luskin-levels of ignorance and incomprehension. Let us now dive deep into Luskin-levels of outright stupidity and dishonesty.
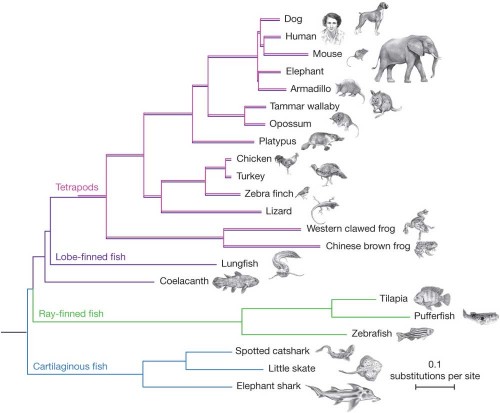
The paper includes this diagram of the evolutionary relationships of various vertebrate species.
These are the relationships determined by comparisons of large portions of the entire genomes of these organisms — it’s a phylogeny based on a large amount of data. Luskin obligingly simplifies it for us:
The closest relative to us tetrapods are lungfish, with coelacanths more distant, and teleosts further still. This is the consensus. It’s supported by many comparisons.
But brace yourself: here’s where Luskin proves that he’s an idiot. He sets aside the synthesis of large data sets, and instead explains that if we look at single genes, evolution is proven wrong!
He read the paper and found a couple of examples interesting genes that were highlighted by the authors themselves. In particular, they cite examples of gene losses. All other vertebrates have a component of the immune system, a gene called immunoglobulin M (IgM), but coelacanths are unusual in lacking this particular gene. There’s another immune system component, IgW, which seems to be a relatively primitive or ancestral immunoglobulin, which we tetrapods lack, and which is also missing in teleosts, but is found in coelacanths and sharks.
So what does Luskin do? He redraws the whole vertebrate phylogeny based on a single gene. He throws away almost the entire data set, and focuses on a single character.
But in an IgW-based tree, tetrapods should be much more closely related to goldfish than to the coelocanth or the lungfish. That’s startling and unexpected — if you’re a proponent of common descent.
Holy crap. That’s simply a lie. No: if you look at individual genes, this is exactly what we expect to see, and we’re completely unsurprised. Every single gene is not expected to change in lockstep with speciation events, and each lineage can experience the loss of subsets of genes independently of the larger clade. What Luskin is claiming is utterly contrary to everything any evolutionary biologist will tell you.
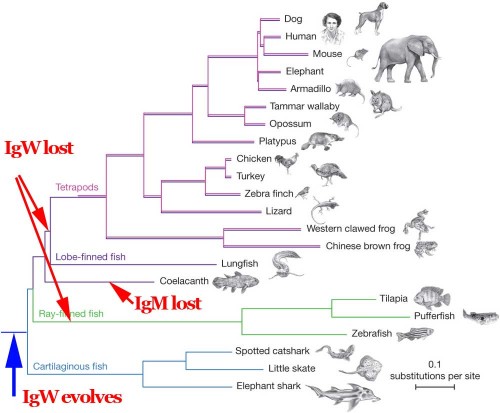
Here’s the original phylogeny from the Amemiya paper. I’ve just added a couple of arrows to indicate simple events in the history of the IgM and IgW genes.
The simplest, most parsimonious explanation is that IgM and IgW were present in the last common ancestor of all of these organisms. The IgW distribution in extant forms is explained by two independent gene losses, one in the ancestor of all tetrapods and another in the ancestor of all teleosts. The IgM distribution is explained by a unique loss in the ancestor of the two modern species of coelacanths. This is trivial and totally unsurprising, requiring no magic, no all-powerful designer, and no processes other than known natural mechanisms of mutation.
Luskin, however, finds this improbable.
Of course this requires some extremely unparsimonious and unlikely events. Since IgW is found in vertebrates as diverse as lungfish and cartilaginous fish (e.g., sharks), a Darwinian evolutionary view would infer that the gene for IgW was present in the ancestor of all jawed vertebrates. If so, then IgW must have stuck around in vertebrates long enough to end up in the sarcopterygian line. But somehow evolutionary theory must explain why tetrapods and all teleosts lack this gene.
“Unparsimonious”? You keep using that word. I don’t think it means what you think it means.
The evolutionary explanation requires three mundane events, the simple loss of a gene in an ancestral population.
The Intelligent Design creationist explanation requires that every extant species was specifically and intentionally stocked with a set of genes hand-chosen by a designer. God magically inserted IgM into each vertebrate species, except that he missed the coelacanths, and he magically inserted IgW into each and every shark, ray, coelacanth, and lungfish, but he intentionally left them out of every tetrapod and teleost.
And Luskin chooses to lecture biologists on the meaning of “parsimonious”? That’s startling, but entirely expected from the frauds at the Discovery Institute.




Luskin: Ha ha, I can’t think, therefore thinking is useless.
Likewise, there was no miracle that caused all genes to change similarly and no miracle preserving IgW in all lineages, therefore a miracle is responsible for all that did happen.
IDiocy, soon to displace Darwinism and all of its evils.
Glen Davidson
I always find it weird when ID proponents both argue that conserved genes are common and argue against common descent.
I had to stop reading partway through (couldn’t take all the idiocy), but it seems like Luskin’s central point is, “No single paper on its own can prove evolution, therefore evolution is not proven.”
Parsimonious, adj, denoting something which can be used to bring more money to the parsonage.
Unparsimonious, adj, denoting something which diverts that money to scientific funding instead.
The gene regulatory region in coelacanth that specifies the formation of lobe fins works in mice. To specify the formation of legs. We already knew the lobe fins and legs were homologous structures but this is another demonstration that we humans are just derived lobe finned fish.
Using that argument, my hair colour would indicate that I am completely unrelated to my brother.
Our mum has got a lot of explaining to do.
(Ah. She’s just reminded me that hair colour depends on two genes, not one. Excuses!)
Does that mean you both are and are not related to your brother, depending on which gene is being examined?
Someone should get Tony Robinson to play Casey Luskin in a movie about the rise and fall of ID. Robinson could recycle his signature line (as Baldrick in the “Blackadder” series): “I have a cunning plan (to defeat evolution).” It would be perfect.
For me, the nadir of this dismal post came in the following passage:
These claims are just utter lies. There’s no other way to describe it. Teleostei is not an “ancient group of vertebrates”; early representatives first appear in the early Triassic, 100 million years after the evolution of the tetrapods. And even if they were an ancient group, the only way that independent loss in tetrapods and teleosts is unlikely is if sarcopterygians evolved from teleosts, which is clearly what Luskin is implying happened. Not only does the timeline not work, but if this were the case then Teleostei would be a paraphyletic group, but the evidence available shows that they are monophyletic. If Luskin is right, then everything we know about the systematics of the major groups of fish is wrong.
So either Luskin has some truly revolutionary evidence up his sleeve or he’s making it up as he goes along. What a conundrum. Gee, I wonder what the answer could possibly be.
At least two. If subtle gradations among basic colors are to be explained, more genes (or at least more alleles) are required.
yeah, evidently Luskin does not grok the difference between teleosts and actinopterygians (the former a subset of the latter).
Casey Luskin’s Posts:
The Grift that keeps on Grifting.
This is a pretty good example of seemingly sophisticated pseudoscience. Anyone with a high school level understanding of biology is going to face palm. They could explain exactly why it is scientifically wrong like PZ can, but they could do a fair rebuttal of it like some of the comments already posted. What is most annoying about DI is that they can fool intelligent but scientifically illiterate people with this crap. In contrast Comfort’s pseudoscience fools mostly derps. That is why it is good to have people, who rebut this for people, who may have simply gotten a deficient science education.
The second most annoying thing about DI is they never conduct their won research. They simply mine other people that do actual scientific research. It is because they can’t generate a testable hypothesis with “God did it”.
Sorry typos. Thinking and typing…
I meant: They couldn’t instead of “They could” and own instead of “won”.
Well, duh. That’s the “heritable” part of descent with heritable modification.
“a Masters degree (in Science! Earth Science, that is)”
Oi, stop that. I’ve got a Masters in Earth Science and there was more than enough phylogeny in my palaeontology courses to know that Luskin is full of shit.
We’re not biologists, but we’re still supposed to know enough biology to avoid being that stupid.
Remember, boys and girls, Casey doesn’t write for competent scientists, he write for creationists who are largely ignorant of science. All he has to do is use “sciency” verbiage and voila! His target audience is thrilled with his brilliant takedown of all of modern biology.
He’s just a useless tool, but a great source amusement for us. Keep on giving, Casey!!
How does this statement even make sense except from an evolutionary perspective?
Seems like it would be nice to sequence some non-teleost ray-finned fish, because they diverged between the divergence of ray-finned and lobe-finned fish and the divergence of various subgroups of teleosts.
Fish like bichirs, sturgeons, gars, and bowfins.
But given how cheap it has become to sequence billion-base-pair genomes, I would not be surprised if at least some of these fish were soon sequenced.
Ha! Merely a flesh wound!
The designer’s perfect indivisible triune nature requires that hox genes are widely conserved among vertebrates.
What unevidenced designer are you talking about?
nope sorry. Your assignment was:
your response still makes no sense, so you fail. :)
could be a trick question though… there is no positive way to answer it.
This is really only a superficially more sophisticated version of the game they play with transitional fossils; they demand evidence, in the form of one single fossil, for evolution, which is a process of gradual change over time in populations, not individuals. Evolution isn’t demonstrated by single fossil examples, it’s seen in lines of fossils that show a process of transition (this is what I’ve always understood “every fossil is a transitional fossil” to mean).
And here’s the funny part- if someone were to one day uncover the “crocoduck” they demand, it wouldn’t, in fact, prove evolution, it would disprove it. They’re demanding to see as “proof” of evolution something they themselves need to produce as evidence for “poof! goddidit!”
When are they going to make the “DI movie”. Where, “Casey get’s out and devices a plan to get rid of science.” And he does it by, “…burning all the records of science, and brainwashing people to creationism.” But, Luskin fails flat on his face, because Dawkin sand the rest of the crew of anti-creationists educate everybody, and stop Luskin from writing books, receiving funds, and broadcasting.
One of the key traits of intelligence is to know enough that you know you don’t know everything, and to understand that everything is contextual, and that you may not have the correct context for the information you have. Luskin fails this simple intelligence test in spades.
*facepalm* Nerd, are you completely unable to read for context? davehooke was mocking Luskin by pretending to answer the question in comment 18, which he quoted in full.
Not sure Hollywood can imagine that.
Stop him from writing books? Only publishers could do that, and that only if he doesn’t self-publish via some book-on-demand service. (…Has he in fact written any book?) Similar for broadcasting. If the Disinformation Institute gets any funds at all, they come from stink tanks…
Just a peripheral comment here. I graduated with a degree in Biology with an emphasis in evolution (some years ago) and although I attempt to keep up with the literature I find it impossible to stay on top. Whenever I learn something new (to me) such as this finding and ponder its implications as well as the research and dedication involved in its discovery all I can do is close my eyes, lean back and say, “Wow”. I let the awe and wonder of life and nature wash over me and marvel at our ability to unravel at least a bit of it. The universe is truly astonishing and wonderful and life is grand.
Funny part is, when someone asked about this over at Rational Skepticism, I posted [url=http://www.rationalskepticism.org/biology/coelacanth-genome-sequenced-t38883.html#p1697485]this[/url]by way of reply, before I was pointed at PZ’s exposition, and although my explanation is probably nowhere near as polished or robust as the above, I appear to have alighted upon the essential details, despite having only elementary knowledge of the workings of molecular phylogeny.
Yet Casey Luskin was either too stupid to deduce this for himself, or deliberately avoided doing so because it would drive a tank division through his assertion-laden, mythology-based presuppositions. Or, possibly, both apply here.
I’m sure PZ will have fun picking holes in my elementary presentation for didactic purposes, but hopefully will tell everyone that whilst I’ve missed out a lot of details, I’m a lot less wrong than Luskin. :)
squueeeeeeek